|
Analyzing and visualizing Next Generation Sequencing data, incorporates cutting-edge technology and algorithms, while also supporting and integrating with the rest of your typical NGS workflow.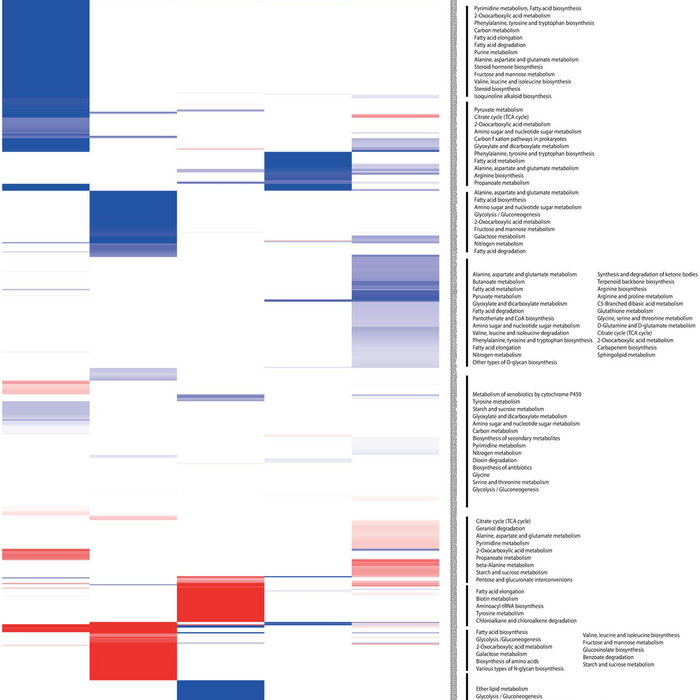 
We will use the pcDNA3-atp8a1 sequence from the Cloning folder in the. In this case specificity is not a big issue: the chance that a primer will bind another part of the (small) plasmid is very small. QIAGEN’s CLC Genomics Workbench helps analyze and visualize next generation sequencing (NGS) data with cutting-edge. We will use the CLC Main Workbench to design primers for checking if a gene is successfully inserted in a vector. Prerequisites To run this tutorial, you must be working with the CLC Genomics Workbench, version 5.5 or higher. This training will be held in the NIH Library Training Room, Building 10, from 9:30 a.m. 4.Import the reference sequence and adapter list provided with the example data by going to: File Import Standard Import Ensure the option Automatic import is selected and select 'MT135044. The NIH Library’s Bioinformatics Support Program has organized a CLC Genomics Workbench class for Wednesday, August 31. Tutorial Bisulfite Sequencing 2 Bisulfite Sequencing This tutorial takes you through a complete bisulfite sequencing workflow using Bisulfite Sequenc-ing plugin for CLC Genomics Worbench 8.5 and higher, and Biomedical Genomics Workbench version 2.5 and higher. 
The CLC Genomics Workbench is the client software for the CLC Genomics Server.įeatures for the Genomics Workbench are described below: 3.Create a new folder for the project with a relevant name, for example, 'SARS-CoV-2 MinION Tutorial'. To download the genome sequences for the Denovo assembly tutorial use the following links, ,, and.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
